Bayesian Model Criticism¶
This tutorial walks through a scoring-and-stacking workflow for
Bayesian models. It demonstrates the features that set trade-study
apart from conventional hyperparameter optimizers: proper scoring
rules, calibration assessment, and score-based model averaging.
The underlying model is a conjugate Bayesian linear regression — no MCMC sampler or external probabilistic programming library needed, only numpy.
Run it yourself
The full runnable script is at
examples/bayesian_study.py.
The problem¶
We want to fit a Bayesian regression \(y = a + bx + \varepsilon\) where \(\varepsilon \sim \mathcal{N}(0,\sigma_\text{true}^2)\) and the true coefficients are known (\(a=2\), \(b=3\), \(\sigma_\text{true}=0.5\)).
The design question is: which prior hyperparameters and sample sizes produce well-calibrated, accurate posteriors? Specifically, each configuration controls three knobs:
| Factor | Meaning |
|---|---|
prior_var |
Variance of the Normal prior on \((a, b)\) |
noise_scale |
Assumed observation noise \(\sigma\) (may disagree with truth) |
n_obs |
Number of training observations |
Choosing noise_scale too small produces overconfident posteriors;
too large gives conservative intervals. A tight prior (prior_var
small) biases the coefficients toward zero. More data always helps,
but at a computational cost.
Ground-truth model¶
# True data-generating process: y = a + b*x + eps, eps ~ N(0, sigma_true^2)
TRUE_A = 2.0 # intercept
TRUE_B = 3.0 # slope
SIGMA_TRUE = 0.5 # observation noise std
N_TEST = 50 # test locations for scoring
RNG_SEED = 42
rng = np.random.default_rng(RNG_SEED)
X_TEST = np.linspace(0.0, 1.0, N_TEST)
Y_TEST = TRUE_A + TRUE_B * X_TEST + rng.normal(0.0, SIGMA_TRUE, N_TEST)
Conjugate posterior¶
Because we use a Normal prior with known noise variance, the posterior over regression coefficients is available in closed form. Given design matrix \(\mathbf{X}\) and prior precision \(\boldsymbol{\Lambda}_0 = \sigma_\text{prior}^{-2}\mathbf{I}\):
We draw posterior predictive samples by projecting coefficient draws through the test design matrix and adding observation noise.
def bayesian_regression(
x_train: np.ndarray,
y_train: np.ndarray,
prior_var: float,
noise_scale: float,
n_samples: int = 500,
seed: int = RNG_SEED,
) -> tuple[np.ndarray, np.ndarray]:
"""Fit a conjugate Bayesian linear regression and draw predictions.
Prior: beta = (a, b) ~ N(0, prior_var * I)
Likelihood: y | x, beta ~ N(X @ beta, noise_scale^2 * I)
Args:
x_train: Training inputs, shape (n,).
y_train: Training targets, shape (n,).
prior_var: Prior variance for each coefficient.
noise_scale: Assumed observation noise standard deviation.
n_samples: Number of posterior predictive draws.
seed: Random seed.
Returns:
Tuple of (posterior_mean_at_test, predictive_samples_at_test).
posterior_mean_at_test has shape (N_TEST,).
predictive_samples_at_test has shape (N_TEST, n_samples).
"""
gen = np.random.default_rng(seed)
# Design matrices
x_design = np.column_stack([np.ones_like(x_train), x_train]) # (n, 2)
x_star = np.column_stack([np.ones(N_TEST), X_TEST]) # (N_TEST, 2)
sigma2 = noise_scale**2
prior_precision = np.eye(2) / prior_var # (2, 2)
# Conjugate posterior: beta | y ~ N(mu_n, Sigma_n)
gram = x_design.T @ x_design # X^T X
posterior_cov = np.linalg.inv(prior_precision + gram / sigma2)
posterior_mean = posterior_cov @ (x_design.T @ y_train / sigma2)
# Posterior predictive at test locations
pred_mean = x_star @ posterior_mean # (N_TEST,)
# Draw beta samples, then project to predictions and add noise
beta_samples = gen.multivariate_normal(
posterior_mean,
posterior_cov,
size=n_samples,
) # (n_samples, 2)
pred_samples = (x_star @ beta_samples.T) + gen.normal(
0.0, noise_scale, (N_TEST, n_samples)
)
return pred_mean, pred_samples
Simulator and scorer¶
The simulator generates training data from the true model, fits the conjugate posterior, and returns posterior predictive samples at held-out test locations. The scorer evaluates three complementary metrics:
- CRPS (Continuous Ranked Probability Score) — a proper scoring rule that rewards sharp, well-calibrated predictive distributions.
- Energy score — a multivariate proper scoring rule complementary to CRPS.
- Coverage (95 %) — the fraction of test points whose true value falls inside the 95 % posterior predictive interval.
- RMSE — root mean squared error of the posterior mean.
- MAE — mean absolute error of the posterior mean.
class BayesianRegressionSimulator:
"""Generate training data and compute the Bayesian posterior.
Each config specifies prior hyperparameters and sample size.
The "truth" is the held-out test set; "observations" are the
posterior predictive samples at those test points.
Args:
n_samples: Number of posterior predictive draws. Fewer draws
give faster but noisier score estimates — useful as a cheap
surrogate in a multi-fidelity workflow.
"""
def __init__(self, n_samples: int = 500) -> None:
"""Initialise the simulator.
Args:
n_samples: Number of posterior predictive draws.
"""
self.n_samples = n_samples
def generate(self, config: dict[str, Any]) -> tuple[Any, Any]:
"""Draw training data, fit posterior, and return test predictions.
Args:
config: Must contain prior_var, noise_scale, n_obs.
Returns:
Tuple of (y_test, observation dict with predictive samples
and posterior mean predictions).
"""
gen = np.random.default_rng(RNG_SEED)
n_obs = round(config["n_obs"])
# Draw training data from the true model
x_train = gen.uniform(0.0, 1.0, n_obs)
y_train = TRUE_A + TRUE_B * x_train + gen.normal(0.0, SIGMA_TRUE, n_obs)
# Compute conjugate posterior
pred_mean, pred_samples = bayesian_regression(
x_train,
y_train,
prior_var=config["prior_var"],
noise_scale=config["noise_scale"],
n_samples=self.n_samples,
)
observations = {
"pred_mean": pred_mean,
"pred_samples": pred_samples,
"n_obs": n_obs,
}
return Y_TEST, observations
class BayesianRegressionScorer:
"""Score posterior predictions with proper scoring rules."""
def score(
self,
truth: Any,
observations: Any,
config: dict[str, Any],
) -> dict[str, float]:
"""Compute CRPS, energy, 95 percent coverage, RMSE, and MAE.
Args:
truth: True test-set values (y_test).
observations: Dict with pred_samples and pred_mean arrays.
config: Hyperparameter dict (unused).
Returns:
Scores for crps, energy, coverage_95, rmse, and mae.
"""
crps_val = score("crps", observations["pred_samples"], truth)
energy_val = score("energy", observations["pred_samples"], truth)
cov95 = score("coverage", observations["pred_samples"], truth, level=0.95)
rmse_val = score("rmse", observations["pred_mean"], truth)
mae_val = score("mae", observations["pred_mean"], truth)
return {
"crps": crps_val,
"energy": energy_val,
"coverage_95": cov95,
"rmse": rmse_val,
"mae": mae_val,
}
Observables and factors¶
Coverage is given lower weight (0.5) because a model can trivially achieve 100 % coverage by being very wide; CRPS already penalises that.
observables = [
Observable("crps", Direction.MINIMIZE),
Observable("energy", Direction.MINIMIZE, weight=0.5),
Observable("coverage_95", Direction.MAXIMIZE, weight=0.5),
Observable("rmse", Direction.MINIMIZE),
Observable("mae", Direction.MINIMIZE, weight=0.3),
]
factors = [
Factor("prior_var", FactorType.CONTINUOUS, bounds=(0.1, 10.0)),
Factor("noise_scale", FactorType.CONTINUOUS, bounds=(0.1, 2.0)),
Factor("n_obs", FactorType.CONTINUOUS, bounds=(10.0, 200.0)),
]
Annotations¶
An Annotation attaches external metadata — here, a simple linear
cost model for the number of observations:
Constraints¶
A Constraint defines a hard feasibility bound on an observable.
feasibility_filter builds a phase filter that drops any design
violating the constraints:
# Require at least 90% empirical coverage at the 95% nominal level
min_coverage = Constraint(
name="min_coverage",
observable="coverage_95",
op=">=",
threshold=0.90,
)
Factor screening¶
Before running the full grid, Morris screening identifies which
factors influence the scores most. reduce_factors drops any
factor whose \(\mu^*\) falls below a threshold:
def run_fn(config: dict[str, Any]) -> dict[str, float]:
"""Compose simulator + scorer for screening.
Returns:
Score dictionary from the scorer.
"""
config = {**config, "n_obs": round(config["n_obs"])}
truth, obs = world.generate(config)
return scorer.score(truth, obs, config)
importance = screen(run_fn, factors, method="morris", n_trajectories=8, seed=42)
print("Morris screening (mu_star):")
for name, vals in importance.items():
print(f" {name}: {vals}")
reduced = reduce_factors(factors, importance, threshold=0.01)
print(f"\nImportant factors (threshold=0.01): {[f.name for f in reduced]}")
Grid evaluation¶
A 60-point Latin hypercube covers the three-factor space. run_grid
evaluates every configuration and collects scores and annotations into
a ResultsTable:
grid = build_grid(factors, method="lhs", n_samples=60, seed=42)
for cfg in grid:
cfg["n_obs"] = round(cfg["n_obs"])
print(f"\nLHS grid: {len(grid)} configurations")
results = run_grid(
world=world,
scorer=scorer,
grid=grid,
observables=observables,
annotations=[compute_cost],
)
Pareto analysis¶
print(f"\nPareto front: {len(front_idx)} / {results.scores.shape[0]} designs")
header = (
f"{'prior_var':>10s} {'noise':>6s} {'n_obs':>5s} "
f"{'CRPS':>6s} {'Energy':>7s} {'Cov95':>6s} "
f"{'RMSE':>6s} {'MAE':>6s} {'Cost':>5s}"
)
print(f"\n{header}")
print("-" * len(header))
for i in front_idx:
cfg = results.configs[i]
crps_val = results.scores[i, 0]
energy_val = results.scores[i, 1]
cov = results.scores[i, 2]
rmse_val = results.scores[i, 3]
mae_val = results.scores[i, 4]
cost = results.annotations[i, 0] if results.annotations is not None else 0.0
print(
f"{cfg['prior_var']:10.2f} {cfg['noise_scale']:6.2f} "
f"{cfg['n_obs']:5d} "
f"{crps_val:6.3f} {energy_val:7.3f} {cov:6.3f} "
f"{rmse_val:6.3f} {mae_val:6.3f} {cost:5.1f}"
)
Pareto front scatter matrix¶
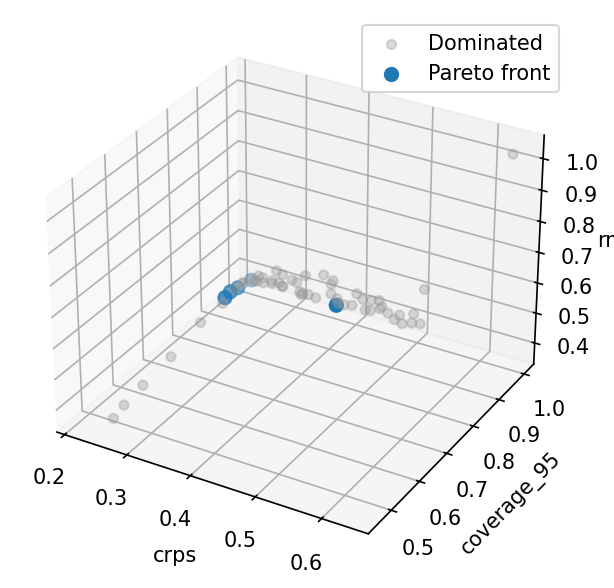
With three objectives the front is a surface; plot_front shows all
pairwise projections. Models with low CRPS generally have good
coverage and low RMSE, but there are trade-offs when noise is
mis-specified.
Parallel coordinates¶
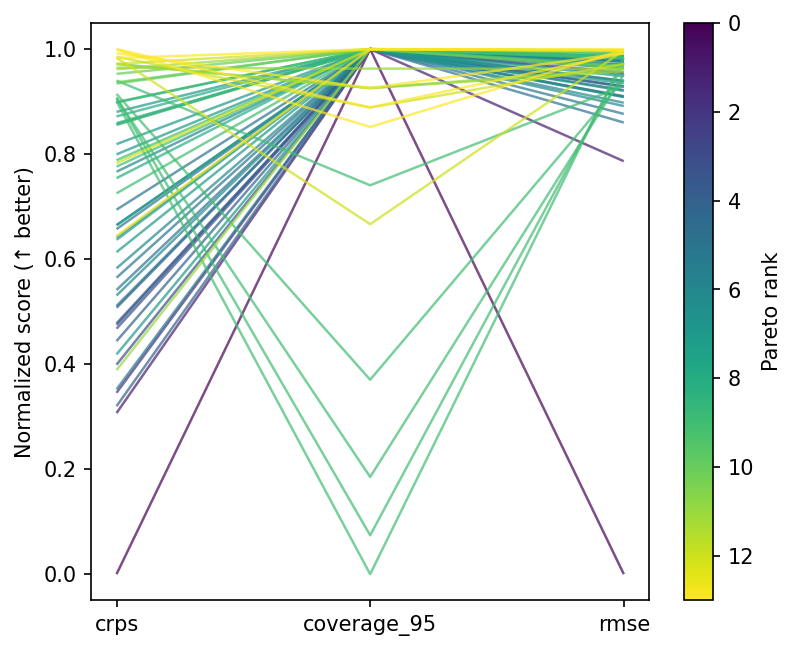
Front designs cluster at moderate prior variance and noise scales close to the true value — the region where the model is neither overconfident nor excessively vague.
CRPS strip plot¶
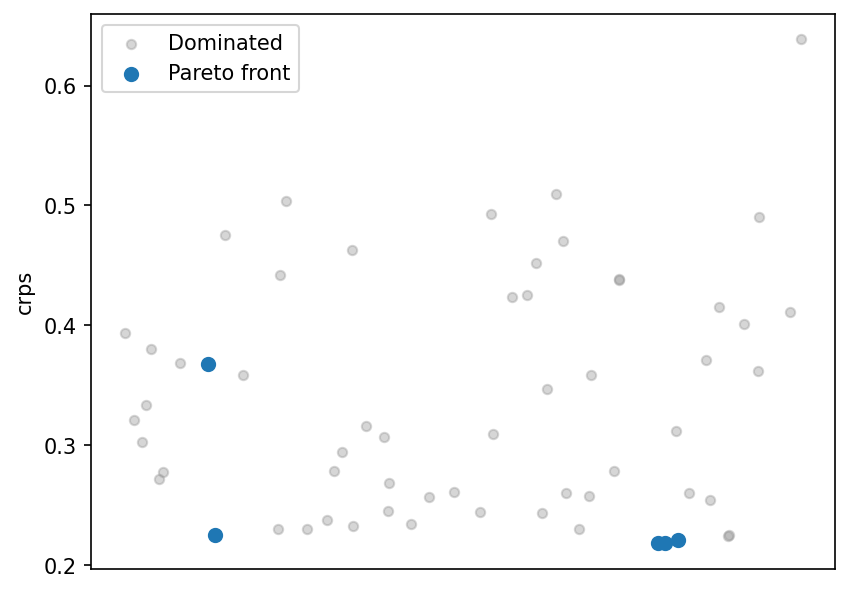
Quality indicators¶
hypervolume measures the dominated volume of the Pareto front;
igd_plus (inverted generational distance plus) measures how close
the front is to a reference set. Together they quantify front
quality — useful for comparing design methods or grid sizes:
directions = [o.direction for o in observables]
weights = [o.weight for o in observables]
front_scores = results.scores[front_idx]
# Build a synthetic reference front from per-objective best values
n_obj = front_scores.shape[1]
ideal = np.tile(np.median(front_scores, axis=0), (n_obj, 1))
for j, d in enumerate(directions):
ideal[j, j] = (
front_scores[:, j].min()
if d == Direction.MINIMIZE
else front_scores[:, j].max()
)
ref_point = np.array([
front_scores[:, j].max()
if d == Direction.MINIMIZE
else front_scores[:, j].min()
for j, d in enumerate(directions)
])
hv = hypervolume(front_scores, ref_point, directions, weights)
igd = igd_plus(front_scores, ideal, directions, weights)
print(f"\nHypervolume: {hv:.4f}")
print(f"IGD+: {igd:.4f}")
Feasibility filtering¶
Apply Constraint + feasibility_filter to keep only designs
meeting hard bounds (here, coverage_95 >= 0.90):
all_idx = np.arange(results.scores.shape[0])
feas_filter = feasibility_filter(constraints=[min_coverage])
kept = feas_filter(results, observables)
print("\nFeasibility filter (coverage_95 >= 0.90):")
print(f" {len(all_idx)} total -> {len(kept)} feasible designs")
Weighted-sum filtering¶
weighted_sum_filter scalarises objectives into a single score
via min-max normalisation and weighted combination, then keeps
the top-K designs. Useful when you want a simple ranking
without full Pareto analysis:
ws_filter = weighted_sum_filter(
weights={"crps": 1.0, "coverage_95": 0.5, "rmse": 0.8},
k=8,
)
kept = ws_filter(results, observables)
print("\nWeighted-sum filter (top 8 by scalarised score):")
for i in kept[:5]:
cfg = results.configs[i]
print(
f" prior_var={cfg['prior_var']:.2f}, "
f"noise={cfg['noise_scale']:.2f}, "
f"n_obs={cfg['n_obs']}"
)
if len(kept) > 5:
print(f" ... and {len(kept) - 5} more")
Sobol grid¶
build_grid(method="sobol") generates quasi-random points with
better space-filling properties than LHS for low dimensions:
sobol_grid = build_grid(factors, method="sobol", n_samples=40, seed=0)
for cfg in sobol_grid:
cfg["n_obs"] = round(cfg["n_obs"])
print(f"\nSobol grid: {len(sobol_grid)} configurations")
sobol_results = run_grid(
world=world,
scorer=scorer,
grid=sobol_grid,
observables=observables,
)
directions = [o.direction for o in observables]
weights = [o.weight for o in observables]
sobol_front = extract_front(sobol_results.scores, directions, weights)
print(f"Sobol Pareto front: {len(sobol_front)} designs")
Study workflow¶
The Study class orchestrates multi-phase workflows.
Study.summary() returns per-phase statistics and
Study.stack() computes stacking weights directly:
world = BayesianRegressionSimulator()
scorer = BayesianRegressionScorer()
grid = build_grid(factors, method="lhs", n_samples=60, seed=42)
for cfg in grid:
cfg["n_obs"] = round(cfg["n_obs"])
study = Study(
world=world,
scorer=scorer,
observables=observables,
phases=[
Phase(
name="explore",
grid=grid,
filter_fn=feasibility_filter(
constraints=[min_coverage],
),
),
Phase(name="rank", grid="carry"),
],
annotations=[compute_cost],
)
study.run()
# Study.summary() — per-phase statistics
summary = study.summary()
for phase_name, stats in summary.items():
print(f"\n Phase '{phase_name}':")
print(f" trials={stats['n_trials']}, front={stats['n_front']}")
for obs_name, rng in stats["observable_ranges"].items():
print(f" {obs_name}: [{rng['min']:.4f}, {rng['max']:.4f}]")
# Study.stack() — convenience method for score-based stacking
stacking_w = study.stack("rank", maximize=False)
print("Study.stack() weights (top entries):")
for i, w in enumerate(stacking_w):
if w > 0.01:
print(f" design {i}: {w:.3f}")
Score-based stacking¶
stack_scores finds simplex-constrained weights that minimise the
composite MSE across test points. The stacked ensemble often beats
every individual model because it smooths over prior mis-specification:
# Build a score matrix from the front: per-test-point squared errors
front_predictions = []
score_matrix_rows = []
for i in front_idx:
cfg = results.configs[i]
_, obs = world.generate(cfg)
front_predictions.append(obs["pred_mean"])
sq_errors = (obs["pred_mean"] - Y_TEST) ** 2
score_matrix_rows.append(sq_errors)
score_mat = np.array(score_matrix_rows) # (n_front, n_test)
stacking_weights = stack_scores(score_mat, maximize=False)
print("\nStacking weights (MSE-optimal):")
for idx, w in zip(front_idx, stacking_weights, strict=True):
if w > 0.01:
cfg = results.configs[idx]
print(
f" prior_var={cfg['prior_var']:.2f}, "
f"noise={cfg['noise_scale']:.2f}, "
f"n_obs={cfg['n_obs']}: w={w:.3f}"
)
ens_mean = ensemble_predict(front_predictions, stacking_weights)
ens_rmse = float(np.sqrt(np.mean((ens_mean - Y_TEST) ** 2)))
best_single = float(results.scores[front_idx, 3].min()) # rmse column
print(f"\nBest single-model RMSE: {best_single:.4f}")
print(f"Stacked ensemble RMSE: {ens_rmse:.4f}")
Calibration assessment¶
coverage_curve evaluates empirical coverage at many nominal levels.
A well-calibrated model tracks the diagonal:
best_crps_idx = front_idx[int(np.argmin(results.scores[front_idx, 0]))]
best_cfg = results.configs[best_crps_idx]
_, best_obs = world.generate(best_cfg)
nominal, empirical = coverage_curve(best_obs["pred_samples"], Y_TEST)
pv = best_cfg["prior_var"]
print(f"\nCalibration (best CRPS, prior_var={pv:.2f}):")
for level in (0.5, 0.9, 0.95):
closest = empirical[np.argmin(np.abs(nominal - level))]
print(f" Nominal {level:.0%} -> Empirical {closest:.1%}")
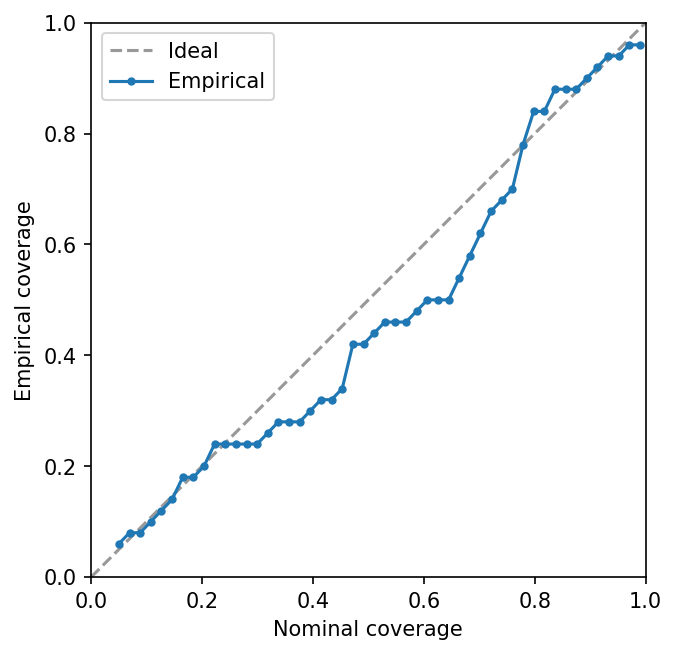
Persistence¶
save_results writes the ResultsTable to disk; load_results
reconstructs it. Useful for expensive studies where you want to
separate evaluation from analysis:
with tempfile.TemporaryDirectory() as tmp:
save_results(results, tmp)
loaded = load_results(tmp)
n_saved = loaded.scores.shape[0]
print(f"\nSaved and reloaded {n_saved} results successfully.")
Multi-fidelity workflow¶
Real-world studies often have an expensive simulator — full MCMC, a fine-mesh CFD solver, or an agent-based epidemiological model. Running every candidate design at full fidelity is wasteful. A multi-fidelity strategy screens many designs cheaply, then validates only the promising ones at high fidelity.
Phase supports this via optional world and scorer overrides.
When set, a phase uses its own simulator instead of the Study-level
default. Here the cheap surrogate draws only 50 posterior samples
(fast but noisy CRPS estimates), while the validation phase draws
2 000:
# Cheap surrogate: only 50 posterior draws (fast, noisier scores)
cheap_world = BayesianRegressionSimulator(n_samples=50)
# Expensive model: 2000 posterior draws (slow, precise scores)
expensive_world = BayesianRegressionSimulator(n_samples=2000)
# Phase-level scorer override: use a separate scorer for screening
screen_scorer = BayesianRegressionScorer()
validate_scorer = BayesianRegressionScorer()
grid = build_grid(factors, method="lhs", n_samples=60, seed=42)
for cfg in grid:
cfg["n_obs"] = round(cfg["n_obs"])
study = Study(
world=expensive_world,
scorer=validate_scorer,
observables=observables,
phases=[
# Phase 1: screen with cheap world AND its own scorer
Phase(
name="screen",
grid=grid,
world=cheap_world,
scorer=screen_scorer,
filter_fn=top_k_pareto_filter(k=10),
),
# Phase 2: validate top 10 with the expensive model + default scorer
Phase(name="validate", grid="carry"),
],
annotations=[compute_cost],
)
study.run()
screen_r = study.results("screen")
validate_r = study.results("validate")
print("\nMulti-fidelity study:")
print(f" Screen phase: {screen_r.scores.shape[0]} designs (50 draws)")
print(f" Validate phase: {validate_r.scores.shape[0]} designs (2000 draws)")
directions = [o.direction for o in observables]
weights = [o.weight for o in observables]
front_idx = extract_front(validate_r.scores, directions, weights)
print(f" Final Pareto front: {len(front_idx)} designs")
The Study orchestrates both phases: the first screens 60 designs
with the cheap surrogate and keeps the top 10 by Pareto rank, then
the second re-evaluates those 10 designs with the expensive model.
This pattern applies whenever fidelity is a computational strategy rather than a design factor:
- Epidemiology — screen surveillance designs with a deterministic ODE model, validate the best with a stochastic agent-based model.
- Engineering — coarse mesh for broad exploration, fine mesh for the Pareto front.
- Forecast model grading — fast approximate inference for screening, full HMC for final assessment.
What to try next¶
- Try
stack_bayesian()on models that expose log-likelihood (requiresarviz). - Add more phases to the
Study— e.g. a third phase that re-evaluates with even higher fidelity. - Experiment with different
weighted_sum_filterweight vectors to explore different trade-off preferences.